Der Gametwist simuliert Ein echtes Online-Casino-Erlebnis . Auf den ersten Blick sieht alles sehr echt aus. Sie können aus Dutzenden von wählen Online-Slots und andere Spiele . Die Spiele werden sogar von einem echten Softwareanbieter übernommen. Das heißt, dass Sie diese Spiele tatsächlich in einigen echten Casinos spielen können.
Das Gametwist Kasino. ist völlig kostenlos. Sie zahlen nichts zum Herunterladen oder Registrieren. Wenn Sie sich nicht für den Kauf von Wendungen entscheiden, werden Sie tatsächlich nichts bezahlen.
Wie spiele ich im Gametwist Online Casino
In Gametwist zu kommen ist viel einfacher als Registrieren Sie sich im Echtgeld Casino . Sie müssen nur einen Spitznamen auswählen, ein Kennwort einrichten, ein Geburtsdatum schreiben und eine gültige E-Mail haben. Sie erhalten 3000 Wendungen zur Bestätigung der E-Mail und Sie können mit dem Wetten zum Spaß beginnen. Sie können Gametwist direkt in Ihrem Browser auf Ihrem Desktop oder Laptop spielen oder eine mobile App auf iOS oder Android herunterladen.
Da kein echtes Geld von Gametwist zurückgezogen werden kann, müssen Sie keine Identitätsnachweise oder andere mit ihm verbundene personenbezogene Daten bereitstellen. Sie bestätigen einfach Ihre E-Mail und spielen Sie. Nach allgemeinen Geschäftsbedingungen ist das Spiel jedoch vorgesehen Für Spieler über 18 Jahre alt . Auch können Minderjährige es nicht spielen.
Als neuer Spieler beginnen Sie auf der ersten Ebene und fahren dann mit jeder Wette vor. Mit jedem Level erhalten Sie eine bestimmte Anzahl von Wendungen als Geschenk. (z. B. auf der zweiten Ebene 3000, in den 3. Stufe 4000 usw.)
Holen Sie sich die Gametwist-App auf Ihr Handy!
Du kannst Laden Sie Gametwist herunter direkt zu Ihnen Android oder iOS-Telefon . Danach können Sie praktisch jederzeit spielen und irgendwo wie Sie möchten. Um die Anwendung verwenden zu können, müssen Sie sich zuerst kurz anmelden.
Wie man Spielen Sie Slots kostenlos Bei Gametwist auf Ihrem Handy:
- Laden Sie die App herunter in: ios gametwist. oder Android Gametwist.
- Holen Sie sich Ihren Willkommensbonus
- Genießen Sie die besten Slots und anderen Casino-Spielen jederzeit, überall!
Welche Casino-Spiele bietet Gametwist an?
Da wir bereits zu Beginn des Gametwist Casino-Überprüfungen erwartet haben, hat ein echter Softwareanbieter in die Anwendung eingefügt. Dies ist der bekannte Anbieter Novomatic. Bzw, Griesenstube und 707 spiele. die beiden unter den Novomatischen fallen.
Es gibt Treffer wie Bizzling Hot, Fruits'n Sevens, Buch von Ra, Golden Sevens, 5 Line Mystery und mehr. Insgesamt können Sie rund 600 Online-Slots an der Gametwist Kasino. .
Gametwist Tipp: Einige Online-Slots bieten höhere Wetten und mehr Erfahrung pro Spin. Mit diesen Spielen ist Ihr Fortschritt schneller.
Die Mindesteinsetzung ist in der Regel auf 200 Wendungen eingestellt einarmige Banditen . Die maximale Wette basiert auf Ihrem Niveau und auf dem Spiel. Am Anfang beträgt jedoch rund 2.500 Wendungen. Es gibt keine Rückkehr zu den erwählten Spielerfiguren, wie dies der Fall ist echte Online Casinos . Wir haben jedoch die Spiele getestet und geschätzt, dass sie etwa 95% verraten sollten.
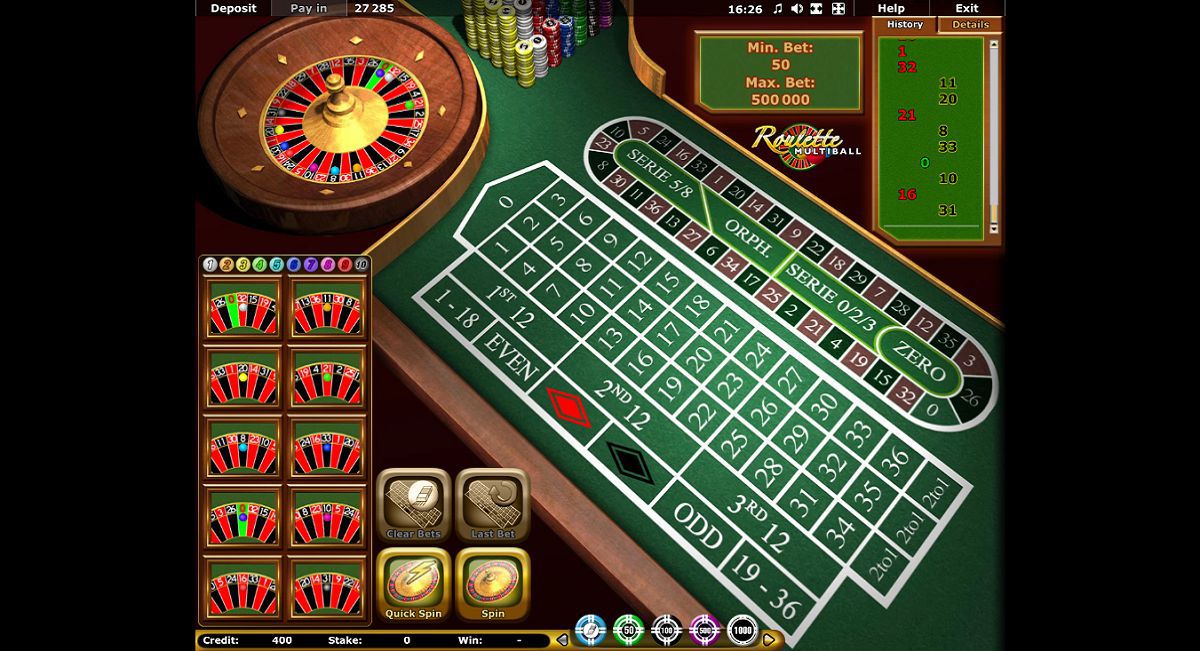
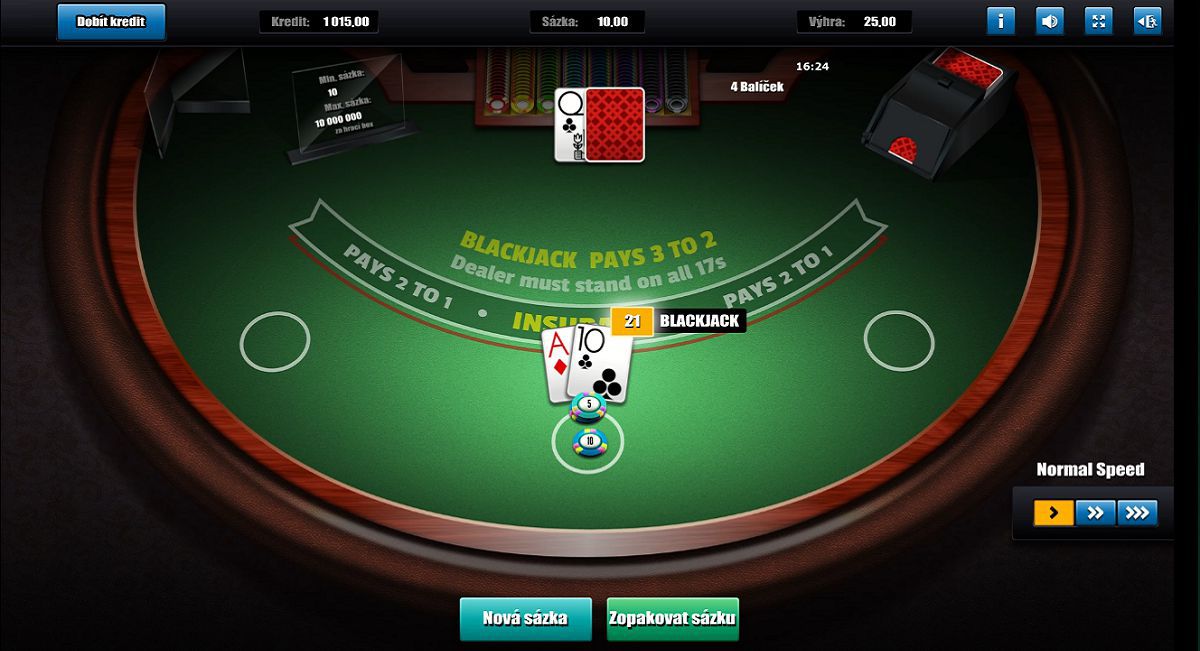
Neben Slot-Maschinen können Sie auch mehrere Versionen von abspielen Roulette, Totschläger. und Baccarat ich bin Gametwist Kasino. Im Poker-Bereich ist eine Hold'em-Version von Poker und mehrere Varianten von Video Poker verfügbar.
Wie bekomme ich einen Gametwist-Bonus?
Um Spiele zu spielen, müssen Sie eine Spielwährung mit dem Namen Wendungen haben. Diese können auf verschiedene Weise erhalten werden. Zuerst bekommst du sie aus dem Spiel:
- Täglicher Anmeldebonus
- Bestätigung - 3.000.
- Zeitbonus (alle zwei Stunden) - auf Ebene (ab 7.000)
- Gewinnende Kasino-Spiele.
- Von Gutscheinen, die per Post gehen
- Twists können auch für echtes Geld im Abschnitt Promotions erworben werden. Wenn Sie jedoch nicht ausgeben möchten, warten Sie einfach und lassen Sie die täglichen und zeitlichen Abläufe beginnen.
Notiz:
- Das Spiel ist für Spieler über 18 Jahre alt
- Spielen spielen kann süchtig machen
- Gewinne können nicht in echtes Geld umgewandelt werden
- Enthält Anzeigen und Einkäufe und Apps


 EIN
EIN
 Tschechisch
Tschechisch
 Polieren
Polieren
 Slowenisch
Slowenisch
 Russisch
Russisch
 Deutsch
Deutsch
 Slowenisch
Slowenisch
 Null
Null
 Schwedisch
Schwedisch
 Portugiesisch
Portugiesisch
 Italienisch
Italienisch
 Spanisch
Spanisch
 Französisch
Französisch
 finnisch
finnisch
 Luke Seidel
Luke Seidel









Du musst sein eingeloggt einen Kommentar hinzufügen